Caroline Chung | Division of Radiation Oncology and Division of Diagnostic Imaging, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA
Citation: EMJ Radiol. 2026; https://doi.org/10.33590/emjradiol/130X4G08
![]()
What first drew you to radiology, in particular the intersection of oncology, imaging, and data science? How has it shaped your career path since?
Clinically, I trained as a radiation oncologist, but since the start of my career I’ve had a faculty role, cross-appointed between radiation oncology and diagnostic imaging (or radiology) due to my research focus on the use of quantitative imaging in personalised cancer therapy. This intersection, together with my training, led me to focus on questions like: can we actually extract quantitative imaging biomarkers or surrogates of tumour biology behaviour response to therapy?
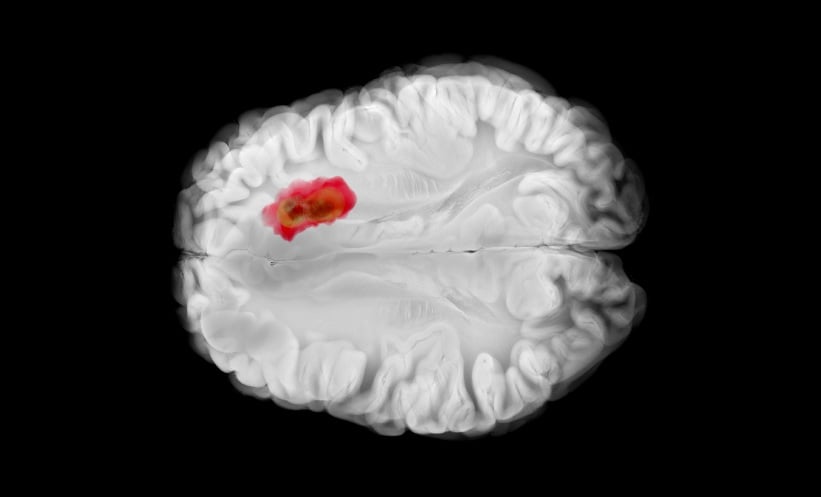
As a radiation oncologist, I was very interested in mapping out resistant or aggressive sub-regions of tumour with imaging, as these are areas could benefit from treatment intensification with locally directed therapy like radiation therapy. At the start of my career, a lot of my work was in that translational space, working right from early mouse models of brain tumours through to clinical trials, and trying to translate those quantitative imaging biomarkers, both from the pre-clinical to clinical space. In recent years, a lot of my focus has been on clinical research development and how we actually translate what is developed in research into clinical practice.
Your clinical practice focuses on malignancies of the central nervous system (CNS). How have advances in imaging, quantitative analysis, and data integration changed the way clinicians detect, characterise, and monitor CNS tumours during your career?
There have been lots of new imaging techniques. In term of MRI, we have identified many novel sequences and various quantitative data we can extract from images, and how have the ability to leverage emerging analytic pipelines (including some using AI) in multi-parametric MRI analysis. Combining MRI with PET and other imaging modalities over the course of time has been something that has also generated new insights in the CNS tumour space. I think what’s most interesting is that each of those modalities brings us different information. What remains unanswered is how do we actually integrate and extract the meaningful pieces of what each of these modalities are contributing? And how do you meaningfully map them together spatially to best represent the underlying biology ?
Another gap is being able to extract quantitative measures and tying them back to the underlying biology. We do need more research in the space. If we can get spatially resolved and temporally resolved biopsy specimens so that we can correlate true pathology to the imaging, we will be closer to understanding what the imaging signals represent biologically. Gathering enough of this kind of information, we can start to really aspire to get in vivo biological understanding from our radiological images, which would be a really helpful tool, particularly in the CNS space, where a biopsy currently means a craniotomy, or some form of invasive access through the skull, into the brain. Having those in vivo imaging biomarkers would be incredibly useful to accelerate our understanding, but also to inform treatment selection and treatment response assessment, particularly as systemic therapies are becoming more and more biologically targeted, as opposed to using generic cytotoxic agents to treat cancer.
You have an extensive and highly influential research portfolio. What key areas is your research currently focusing on? What results are you expecting, and how do you see this work translating into meaningful improvements in patient outcomes and clinical decision making?
I think I’ll start with the big picture and then break it down to some of the fundamental, foundational gaps that we continue to have; gaps that we’re also trying to tackle with a lot of the computational capabilities that are emerging, including AI and physics-informed models.
There’s real aspiration in terms of the personalised medicine space to build digital twins that will journey along with that patient. And it doesn’t necessarily mean an entire human being digital twin, although that is an aspirational blue-skies goal in the field. It could be that, for a patient with cancer, we could build a model of their particular tumour and the surrounding biology so we can anticipate or model and simulate how different treatments might work and see which would result in the best possible outcome for that patient, which would reduce the kinds of toxicities that patient might experience. Building in these aspects would help us move toward truly data-informed, personalised decision-making processes above and beyond what we have today. Currently, evidence-based medicine relies on the results from trials and data that reflect population averages.
Moving into some gaps that we currently have in the imaging space, in order to measure biological changes across time in the imaging, we need to consider the level of uncertainty or variability of the imaging measurement from scan to scan. People go to different scanners or different institutions, which may have different protocols; not all patients stay in one hospital for the entirety of their care. Some of the fundamental work that I’ve been pursuing with others in terms of the quantitative imaging is figuring out a way to meaningfully cross-calibrate these differences to determine true biological change. We know there’s going to be some level of heterogeneity because machines will always get upgraded, and it is great to have technology advance over time, but how do you calibrate the measurement at one time point versus another? I serve as the Co-president for the Quantitative Medical Imaging Coalition (QMIC),1 which is a non-profit organisation that aims to serve as a trusted third party across a growing network of dedicated imaging experts, imaging physics experts, academic institutions and organisation, industry partners, etc., who are working to help enable quantitative imaging practices. This community will need to work together to facilitate the realisation of the full potent emerging technologies like digital twins, because patients will travel between institutions. It will need to be an international effort, and the problem is global when it comes to creating quantitative measurements.
The second point is that if we want to leverage the growing capabilities of AI, it’s sensitive to the data itself. I’m sure everyone’s aware of the phrase ‘garbage in, garbage out’. But what that really means from an imaging standpoint is we need to know how much uncertainty you have in your measurement, which will also depend on how your images are acquired, and if you are looking to measure a change across time, the differences between time point A and at time point B matter. Otherwise, we wouldn’t know if something is a true biological signal or just a detection of the technical noise. What we’re after in terms of personalised medicine is deciphering and teasing out the biological signal and the change that’s happening to inform clinical decisions, while figuring out how to alleviate or account for all of the technical noise that comes with real-world data.
Through initiatives such as the Tumor Measurement Initiative (TMI) and your leadership at the Institute for Data Science in Oncology, you’ve helped build infrastructure to enable clinically meaningful use of data. For quantitative imaging to truly support precision medicine, particularly in the context of tumour heterogeneity, reproducibility and standardisation are critical. What role do coalitions and collaborative frameworks, such as the Quantitative Imaging Biomarkers Alliance, play in making quantitative imaging clinically reliable? And what steps should institutions take to align with these standards and realise the full value of a quantitative approach?
Imaging data have been treated as ‘images’ for many years, born out of initial film-based radiology, but the digital capture of medical imaging data brings forward many opportunities to treat imaging data as ‘assays’ of measurements. This shift in perspective introduces the need for changes in radiology practices and workflows that would also bring a lot of benefits in regards to efficiency, quality, and enabling emerging technology like AI. Within TMI, we aimed to consider the entire imaging chain from image acquisition, image processing, and analysis to improve the consistency, precision, and transparency across this process.
While the quantitative approach is not necessarily all about standardisation, as technology and practices will evolve and advance over time, some level of standardisation is helpful, and it is more important to standardise how we collect the information, so that we know how to cross-calibrate the data even if they are different, whether it’s imaging data or any other data. If we collect enough metadata and have enough descriptors around how that data was captured, we can at least start to understand how data at one time point is different from another time point.
I refer to finding and using ‘fit-for-purpose’ data. Sometimes you can work with noisy data, depending on your purpose. If you’re just looking at general patterns or trends, you can potentially work with noisy data in that exploratory space. But if you’re trying to make a very specific decision about, let’s say, a clinical decision around a patient, you would want to have more precise and less noisy data to have greater confidence. Right now, in medicine, a lot of the work is still fairly manual. Humans are nearly always in the loop. But as we start to introduce more tools along this process, it may not be so transparent to the human in the loop what assumptions have been made, or what uncertainties are being integrated into the outputs they’re looking at and using as they make decisions.
As we start to think about how we’re going to use these tools, there needs to be a collaborative endorsement and adoption of a certain practice of how data is captured, as well as communicated from one group to another. That coordination really relies on coalitions, communities, and even perhaps, to some level, regulatory bodies that would actually help bring forward and highlight the importance. Some of that may come through policy changes, but it also requires awareness and a willingness to adapt, because it ultimately benefits all of these different communities if we move toward a truly coordinated health system. From a data perspective, I would say we still have a long way to go.
I also think that there has been a lot of effort on the technical interoperability. The Digital Imaging and Communications in Medicine (DICOM) standards have helped allow imaging data in digital form to move from one centre to another. That’s step one, which we’re improving at. The next step is that we need the understanding of how imaging data set A differs from imaging data set B. For instance, if we’re taking measurements from each, how do we account for that difference in how the images were acquired and processed? That will be impacted by details such as what tracer or contrast agent was used, how it was injected, as well as all of the imaging parameters for the acquisition that is captured by the scanner.
Not all of these details necessarily require more work to capture. It’s about allowing the metadata to flow and having people recognise that that metadata is really important. Achieving these practices will bring forward the transparency in the data needed for meaningful calibration of the measurements.
While AI tools are increasingly being introduced into clinical workflows, concerns remain around reliability, bias, and real-world performance. What practical guidance would you offer clinicians to help them safely evaluate, implement, and monitor these tools? And how might this ultimately improve patient care?
The field is moving very quickly, and the regulatory bodies are trying their best to keep up and create established benchmarks that are needed for regulatory approval as best as they can. What’s becoming apparent is that getting through regulatory approval is really only step one. Whether it works in your environment or not will depend on whether the training and test data set that was used for regulatory approval looks anything like your data at any given time.
This could mean variations in anything from the types of diseases that are more predominant in your area, to your local demographic makeup in terms of your patient population. It may also depend on the technical aspects of the scanners and protocols you are using. So there’s the human biology side, and the technical side. If the technical nuances of how you’re acquiring your images are very different from what was used in your training and test set, this model may not perform well in your environment.
Because of this, there’s a growing recognition that local validation is really important to understand whether a model works in your particular setting, regardless of whether it has regulatory approval. In essence, FDA approval does not mean it works everywhere by default. So, you do need to test locally (this may be more readily feasible at larger centres and systems at this time because they have the resources to do this). Figuring out how to facilitate and enable all clinics to be able to carry out local validation is something that is still being worked on, and how to do that in the most efficient way is something that is not solved yet.
I would argue that some of the pieces I mentioned earlier (having enough metadata around the data, and having the groups developing these models provide clear descriptions of their data) would help centres at least cross-compare. That could help alleviate the need for a thousand different validation exercises. There are a lot of models out there now, especially in the radiology space, and you don’t want to be testing and validating every single one. You want to identify the ones that are most likely to be relevant to you, and that will be assisted by greater transparency.
There have been discussions around what people call the metadata around the models and descriptions of the model itself. I think it would also help to have additional descriptions around the data used to develop that model. Not revealing the dataset itself, but describing how the data were acquired, the metadata around that data. That would at least allow centres to ask, at a surface level, whether it’s worth going through a full validation exercise. That kind of approach could start to bring some efficiency.
Beyond the initial validation and implementation, ongoing monitoring of model performance is also a critical step, as the data can change across the model lifecycle.
You’ve previously highlighted the concept of ‘digital toxicity,’ where digital tools can unintentionally increase the burden for patients. How can clinicians and healthcare systems balance the benefits of digital technologies with the need to minimise burden and protect patient wellbeing?
It’s definitely a concept we need to keep in mind as we continue to introduce digital tools into healthcare. At my organisation, we wrote an article on digital toxicity to bring awareness of the term to the oncology community,2 and we’ve been proactive about working to coordinate patient-reported outcomes, surveys, other patients engagements across the institution, and gathering feedback; it’s all very important. For instance, someone with cancer will see multiple specialists, and if the practices across specialties are not coordinating with each other, it’s highly possible that same patient could be asked to fill out the same, if not similar surveys over a very short period of time.
We have met to review what surveys are going out for research and for clinical practice. How do we proactively mitigate redundant questionnaires that ask the same or very similar questions? This is a proactive activity that doesn’t actually cost a lot of money; it’s all about coordination. And I think that this is something we do need to really think about, not only across organisations, but across the system. It’s more challenging at the system level, but from a patient perspective, I’m sure that they would appreciate not having to fill out the same forms multiple times.
The other aspect of digital toxicity is that many of the tools we use now could be improved simply by coordinating the flow of data more effectively. If systems spoke to each other better, people might not need to repeatedly enter the same information, even sometimes something as basic as demographic details. Other sectors, particularly marketing, have become very good at this. In healthcare, however, there are important privacy considerations that we have to work through carefully, and that is partly why some of this repetition is very intentional.
It is a fine balance. We need to be confident that we have the right person and the right information, and that records are not being duplicated. In organisations with large volumes of patients, it is entirely possible to have patients with exactly the same first and last name and even possibly the same year of birth. In those cases, repeated clarifying questions can serve a clear safety purpose. The challenge is finding the right balance between repeated digital interactions with questions, surveys, forms, etc., and ensuring that processes are still convenient for patients while remaining safe and accurate.
Another area where this balance comes into play is in the administrative burden that patients increasingly navigate in digital systems. There is a lot of discussion around how emerging tools might help alleviate that burden. In the USA, for example, this often relates to insurance approvals and similar processes. The question is how we can use new technologies to gather the necessary information in a way that is simpler and more intuitive, so that patients do not have to work their way through dense legal language just to understand what information is required to secure approval for treatment.
You are a strong advocate for bringing together a network that cuts across clinical, research, administrative, education, industry, and STEM to bring forward mentorship, sponsorship, and lifelong learning to grow leadership across the cancer space, including through your role as Chair of Women in Cancer-All in Cancer. How has this work contributed to meaningful change, and what more can institutions and leaders do to support the next generation of clinicians and researchers?
We call it Women in Cancer-All in Cancer, because we started out being a women’s physicians organisation, and we actually grew to be very multidisciplinary and interdisciplinary, encompassing everyone. This is because people who weren’t women were saying: “Why can’t I access these resources? This is very helpful.” And I said, “Of course, all should have access to this.” Currently we have a series that was intended to be just a 1-year series, but we’ve continued it on due to its popularity, and it’s called “strengthening through perspectives.” If you really pause and think about that phrase, it shows that the professional challenges, the relationship challenges, and individuals’ internal challenges, whether it be imposter syndrome, negotiation, finding one’s voice or purpose or something else, are common to everybody. Everyone experiences them, but they may not realise others are on that same journey, and yet different areas and different fields or different domains of work will address it in different ways, and we can always learn from each other’s mistakes as well as successes. Also, the time to really think through and invest in personal and professional growth is limited, so finding ways to hear from others from across the field is going to be really important.
My second point is that medicine is no longer hanging up your shackle on a wall and opening up a clinic by yourself. Medicine is growing increasingly interdisciplinary with the integration of technology. There’s an increasing number of people in STEM who are coming to support medicine, people who are very dedicated and just as passionate about that patient outcome and making things better. Sharing perspectives across fields will be key to working effectively together.
An example is that public–private partnerships and industry engagement are something that is clearly necessary for us within academia. We can make great discoveries, but we’re not built to disseminate this out into the public across multiple markets; we have to partner with industry to actually make that impact happen. It needs to be coming from across fields. As the chair of WinC-AlinC, I would encourage any organisation to reach out and engage with us. We’re very open to networking and partnering. For example, we have partnered with multiple organisations across radiation oncology to hold a joint hybrid meeting at the American Society for Radiation Oncology (ASTRO) Annual Meeting for the past 2 years, and we’ve had other ones aligned with various cancer meetings including the American Society for Clinical Oncology (ASCO) and the Canadian Clinical Trials Group (CCTG). We’ve had interest from people outside of cancer, although we are called Women in Cancer-All in Cancer, because I think that everyone is realising that the resources and the approaches that we’re taking from interactive workshops, fireside chats, and dynamic panels to leadership training is something that is applicable to their domains, regardless of where they are.
Looking ahead, what excites you most about the future of radiation oncology and quantitative imaging, and how do you see data enabling more personalised, precise, and equitable care over the next 5–10 years?
I think that integrating the quantitative imaging aspects, teasing out in vivo biology, and really pushing the breadth of what we can manipulate and achieve with radiation delivery could be very transformative. We’re just scratching the surface of all that we can do with how we’re actually delivering radiation.
Today, we’ve become very good at delivering radiation with a high degree of precision to exactly where we want it and currently it is largely delivered with what we call standard fractionation, hypofractionation, and stereotactic radiosurgery schedules. But there are emerging approach such as FLASH radiotherapy, delivering ultra-high doses of radiation at super speeds, and lattice radiotherapy, delivering a mesh where some areas are treated and others are intentionally spared to have different effects on the tumour, normal tissues, and immune response. These capabilities simply weren’t technically feasible if you rewound 15 years, but they are possible today.
Beyond the physical rate and shape of the delivery, there are still many other ways to affect the biology. How many ways can we actually modulate the dose of radiation each day or each week to certain areas while avoiding others? What portions should we be sparing? Where are the immune cells sitting or circulating? There’s a lot of potential to unpack there, and I think that’s really exciting. Once we start thinking about individualising that for each patient, feeding in and calibrating models to individual patient data, we’re no longer saying this is a protocol that everyone goes through. We’re taking the data from one patient, and fine-tuning that radiation treatment plan biologically, not just spatially, where we’re targeting the tumour, but biologically manipulating how we’re delivering that radiation to maximise the outcomes in terms of controlling or eradicating the tumour and minimising the side effects. We’re just getting started making dramatic changes in the outcomes of radiotherapy. Not that they’re not already exciting, but I think we’re just getting started. Now is really an exciting time.