INTRODUCTION
Demodex mites are microscopic arthropods that present in the pilosebaceous units of the human skin.1,2 Demodex mites support the severity of the inflammatory process in dermatoses such as acne, rosacea, seborrheic dermatitis, perioral dermatitis, and can also cause an independent disease called demodicosis.3 Among different methods for the detection of Demodex mites, one of the easiest and most informative is a standardised skin surface biopsy (SSSB).4,5 In practical medicine, it is relevant to search for informative, high-tech, and noninvasive diagnostic methods. Demodex mites can also be detected by in vivo confocal laser scanning microscopy (CLSM) on the facial skin,6 as well as in the terminal bulbs of the eyelashes in ocular demodicosis diagnosis.7 The aim of the study was to compare and analyse the data of in vivo CLSM with SSSB in rosacea patients for the identification of Demodex mites.
MATERIALS AND METHODS
The study recruited 30 participants (12 males [40%] and 18 females [60%]) >18 years old with a diagnosis of rosacea. All patients undertook a SSSB to detect for the presence of Demodex mites and CLSM (VivaScope 1500® Lucid Inc., Rochester, New York, USA). A SSSB was performed in a 1 cm2 area and studied under light microscopy with a magnification of ×40 and ×100. CLSM was carried out at three positions (both cheeks and forehead), in two modes of operation of the VivaBlock and VivaStack. The results of examinations were considered positive if >5 Demodex mites per 1 cm2 were present.
RESULTS
The Demodex mites were found in 24 patients (80.0%) by the SSSB method; 10 patients (33.3%) had <5 mites per 1 cm2 (2.3±1.2 mites on the average). Fourteen patients (46.7%) had >5 mites per 1 cm2, which makes it possible to diagnose demodicosis. The Demodex mites were detected in all patients by CLSM. The mean number of mites found in the follicle was 5.4±3.9 (minimum:1.0; maximum:16.0). Demodex mites were defined as round or long conical formations in the hair follicles, as well as excretory ducts of the sebaceous glands with peripheral hyper contouring (Figure 1).
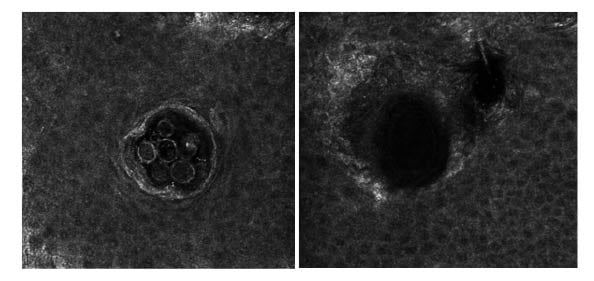
Figure 1: Images obtained with a VivaScope 1500® confocal laser scanning in vivo microscope.
Hair follicles and sebaceous glands with the presence (left) and the absence (right) of Demodex mites.
CONCLUSION
CLSM is an alternative research method for Demodex mite detection, performed noninvasively in a real-time system. This method makes it possible to detect mites in deeper parts of the sebaceous glands that are inaccessible to SSSB. This makes it possible to count the number of mites not only per unit area but also directly in the follicles themselves, and also to analyse the condition of the surrounding skin, underlying structures, and pathomorphological changes.