Presenter: Margaret H. Collins1,2
1. Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA
2. University of Cincinnati College of Medicine, Cincinnati, Ohio, USA
Disclosure: Dr Collins has received consultation fees from Allakos, AstraZeneca, EsoCap, GlaxoSmithKline, Regeneron Pharmaceuticals, Inc., and Shire; and research funding from Receptos, Bristol Myers Squibb, Regeneron Pharmaceuticals, Inc., and Shire.
Acknowledgements: Medical writing assistance was provided by Dr Jennifer L. F. Port, Excerpta Medica, London, UK, funded by Sanofi and Regeneron Pharmaceuticals, Inc.
Support: The publication of this article was funded by Sanofi and Regeneron Pharmaceuticals, Inc.
Citation: EMJ Gastroenterol. 2020;9[1]:41-43. https://doi.org/10.33590/emjgastroenterol/201208
Summary:
Dupilumab is a monoclonal antibody that inhibits IL-4 and IL-13 signalling in multiple Type 2 inflammatory disorders, including eosinophilic oesophagitis (EoE). This article reviews the oral presentation given by Dr Collins at the United European Gastroenterology (UEG) Week Virtual 2020 and describes the results of a post hoc analysis of a Phase II proof-of-concept study of dupilumab in adults with active EoE. The aim of the analysis was to ascertain whether there were any correlations between gene expression and disease severity in patients enrolled in the study.
Background
EoE is a chronic inflammatory disease triggered by an abnormal Type 2 inflammatory response to allergens, and is characterised by eosinophilic inflammation of the oesophagus and histological abnormalities.1 Patients with EoE have an altered oesophageal transcriptome compared with healthy controls, including dysregulation of genes associated with epithelial barrier function and proliferation, collagen and fibrosis pathways, and Type 2 inflammation.2
IL-4 and IL-13 are key and central drivers of Type 2 inflammation in multiple diseases, including EoE. Dupilumab is a fully human monoclonal antibody that binds specifically to IL-4Rα, the shared receptor component of IL-4 and IL-13, and thus inhibits the dual signalling pathways of both cytokines.3-6
Materials and Methods
In a double-blind, placebo-controlled, Phase II proof-of-concept study, adults with active EoE were randomised 1:1 to receive 12 weeks of subcutaneous dupilumab 300 mg weekly or placebo.7 Dupilumab demonstrated efficacy in reducing dysphagia, histological severity (as assessed by EoE histological scores and peak oesophageal intraepithelial eosinophil count), and endoscopic features of mucosal inflammation and remodelling (as assessed by the EoE endoscopic reference score [EoE-EREFS]), and had an acceptable safety profile. In this post hoc analysis of the study, the authors investigated the correlation between gene expression and both histological severity and the macroscopic features of mucosal inflammation and remodelling in 41 of the patients enrolled in the study.
Pinch biopsies were collected from the proximal, mid, and distal oesophagus at baseline and Week 12, and RNA was extracted for transcriptome analysis. Gene expression per patient was averaged across the three samples at each time point. The top 50 most upregulated and top 50 most downregulated genes in EoE that were normalised by dupilumab treatment were used to create a normalised enrichment score (the DpxOme-EoE™ NES). The NES allowed conversion of the expression level of multiple genes to one score that reflected the overall molecular phenotype of a sample. Spearman correlation analysis was performed to compare the DpxOme-EoE NES or individual gene expression with total histological scoring system (HSS) scores, tissue eosinophil count, and EoE-EREFS. The EoE-HSS assessed the severity of histological changes in the oesophagus (grade score) and the extent of tissue that was abnormal (stage score). The EoE-EREFS measured the severity of endoscopic findings.
Results
Dupilumab normalised the expression of 1,302 genes after 12 weeks of treatment, whereas no significant changes were seen in the placebo group. In the dupilumab group, this normalisation resulted in a transcriptome similar to that seen in healthy controls; the transcriptome in the placebo group remained similar to published transcriptome data from patients with EoE.2
Spearman analysis showed strong, significant (all p<0.001) correlations between the DpxOme-EoE NES and the total EoE-HSS grade score (correlation coefficient: 0.832), the EoE-HSS stage score (correlation coefficient: 0.787), and peak oesophageal intraepithelial eosinophil counts (correlation coefficient: 0.773) (Table 1). A moderate correlation (correlation coefficient: 0.562; p<0.001) was observed with EoE-EREFS. A high correlation was found with individual genes, including those related to epithelial proliferation, such as ANO-1; those related to barrier function, such as SPINK8; those related to inflammatory mechanisms, such as CTSC and CRISP3 (coding for the proinflammatory protease cathepsin C and for a cysteine-rich secretory protein, respectively); and GPR97,8 a gene highly expressed on airway eosinophils following allergen challenge.
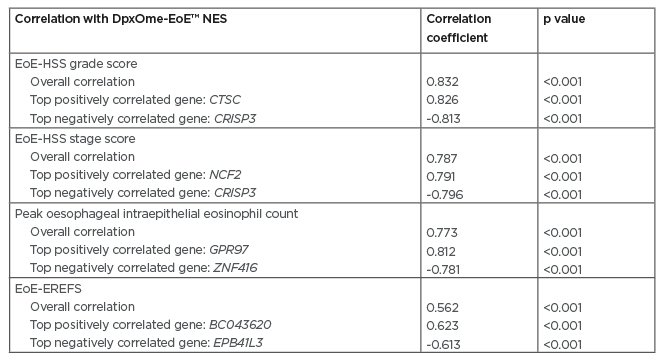
Table 1: Correlations between the post-dupilumab transcriptome and histological and endoscopic endpoints.
DpxOme-EoE™ NES: normalised enrichment score of the top 50 most upregulated and top 50 most downregulated genes in eosinophilic oesophagitis that were normalised by dupilumab treatment; EoE-EREFS: eosinophilic oesophagitis endoscopic reference score; EoE-HSS: eosinophilic oesophagitis histological scoring system.
Conclusion
These data demonstrate that dupilumab normalises several pathways known to be dysregulated in EoE. The post-dupilumab treatment transcriptome in patients with EoE, i.e., the DpxOme-EoE NES, was strongly correlated with EoE-HSS grade score, EoE-HSS stage score, and peak oesophageal intraepithelial eosinophil count, and was moderately correlated with EoE-EREFS. Strong correlations were seen for individual genes associated with epithelial proliferation, barrier function, and inflammatory mechanisms. The expression of genes associated with remodelling in EoE were also among those normalised by dupilumab. Additional analyses are ongoing to further evaluate intrapatient variability. This present analysis suggests that the improvements in histological and endoscopic measures that occur following dupilumab treatment in patients with EoE are due, at least in part, to direct effects on oesophageal epithelial gene expression.