Abstract
Artificial intelligence (AI) is defined as “the ability of a digital machine or computer to accomplish tasks that traditionally have required human intelligence.” Simply put, these are machines that can think and learn. AI has become ubiquitous and is now being applied in healthcare. It uses concepts from statistics, computer science, and data science. An increased understanding of the tools is essential for medical practitioners and medical researchers so that they can contribute meaningfully to the development of the technology, instead of being mere data providers or data labellers. AI has been employed in the finance, marketing, and travel industries for a considerable length of time, but its application in medicine is only gaining acceptance now. Practical office tools based on the AI technologies appear to still be some time away, but these AI-based applications have the potential to benefit all stakeholders in the healthcare industry. This article briefly overviews the current paradigms of AI in medicine.
INTRODUCTION
The Merriam-Webster dictionary defines artificial intelligence (AI) as “a branch of computer science dealing with the simulation of intelligent behavior in computers” or “the capability of a machine to imitate intelligent human behavior.” The layman may think of AI as mere algorithms and programs; however, there is a distinct difference from the usual programs which are task-specific and written to perform repetitive tasks. Machine learning (ML) refers to a computing machine or system’s ability to teach or improve itself using experience without explicit programming for each improvement, using methods of forward chaining of algorithms derived from backward chaining of algorithm deduction from data. Deep learning is a subsection within ML focussed on using artificial neural networks to address highly abstract problems;1 however, this is still a primitive form of AI. When fully developed, it will be capable of sentient and recursive or iterative self-improvement.
Classification
AI and AI-enabled machines can be classified depending on the emulation of the human mind. Technically, AI can be differentiated into three broad types: artificial narrow intelligence (ANI), artificial general intelligence (AGI), and artificial super intelligence.
AGI describes machinery that can work like the human brain and has the capacity to understand or learn any intellectual task that a human can and to transfer this knowledge to other related domains. AGI is also referred to as strong AI, full AI, or as having the ability to perform ‘general intelligent action’.2,3 This is differently viewed by some authors who feel that strong AI is one that is capable of experiencing consciousness;3 however, this is a powerful feat that we are some way away from yet.
ANI, also known as narrow-AI or weak-AI, is the only AI in existence at the present time. It possesses a narrow range of abilities and is focussed on a single, narrow task. Most people are aware of machine intelligence using natural-language-processing (NLP) such as Siri, Cortana, Google Assistant, Google Translate, and Alexa. Some AI systems in medicine diagnose and treat cancers and other illnesses using human-like reason and cognition, and as such are further examples of ANI.
Futuristic artificial super intelligence could in time surpass humans in certain domains because it can not only be applicable on a programmable level, but also in artistic creativity, emotional relationships, and decisions. Although these are strictly human domains, even today the population can struggle at managing emotional relationships.2,3
Current Sources of Data
The main areas in which AI is being applied in healthcare include mass screening, diagnostic imaging, laboratory data, electrodiagnosis, genetic diagnosis, clinical data, operation notes, electronic health records, and records from wearable devices.
Until recently, the most commonly used data in healthcare transactions and research were relational and heterogeneous databases. With the continual integration of the internet of things and advent of wearable technologies, the volume of data available for analysis is increasing and the techniques will also undergo paradigm shifts. This article focusses on the current paradigms and leaves the future direction of AI open to scientific advancements. Formerly, AI techniques used for structured data were ML methods such as classical support vector machines and neural network use, whereas for unstructured data, modern deep learning and NLP were commonly used. Various tools used in oncology, neurology, radiology, ophthalmology, and cardiology have become available.4 Early detection, diagnosis, treatment, prognosis evaluation, and outcome prediction have all benefitted from AI systems, for example, IBM Watson.5
Currently, the use of IBM’s Watson Oncology computer system at the Memorial Sloan Kettering Cancer Center, Manhattan, New York, USA, and the Cleveland Clinic, Cleveland, Ohio, USA, has identified drugs for the treatment of cancer patients with equal or better efficiency than human experts. It can also analyse journal articles for insights on drug development in addition to tailoring therapy.5 At the Oregon Health & Science University (OHSU) Knight Cancer Institute, Portland, Oregon, USA, Microsoft’s Hanover Project has analysed medical research and has predicted the most effective cancer drug treatment option tailored to individual patients with equal efficiency as a human expert.6 The UK NHS has been using Google’s DeepMind platform to detect certain health risks by analysing data that was collected using a mobile phone app. It also analyses medical images collected from NHS patients with the aim to generate computer vision algorithms to detect cancerous cells.7 At Stanford University, Stanford, California, USA, the radiology algorithm performed better than human radiologists in identifying pneumonia, while for meeting the challenges of diabetic retinopathy (DR) management, the computer was as good as the expert ophthalmologists in making a referral decision. In addition, these computers could be on duty all year round, day and night, unlike the human experts.8
MACHINE LEARNING, DEEP LEARNING, AND NATURAL LANGUAGE PROCESSING
AI uses the ML component for handling structured data (images, electrophysiology [EP] data, genetic data) and the NLP component for mining unstructured texts. The algorithms are then trained using healthcare data, meaning the computers can aid physicians with disease diagnosis and treatment suggestions. The IBM Watson system is a pioneering system that has ML and NLP modules, and its application in the oncology field has received much praise. From Watson cancer treatment recommendations, 99% have correlated with decisions of the treating physician.5,9 Watson in collaboration with Quest Diagnostics offer AI genetic diagnostic analysis.10 By analysing genetic data, Watson has successfully identified the rare secondary leukaemia caused by myelodysplastic syndromes in Japan.10
For this discussion, AI devices are divided into two categories: ML techniques, which analyse structured data such as imaging, genetics, and EP data, and NLP methods, which extract information from unstructured data including clinical notes, medical journals, and other sources (Figure 1).4,11
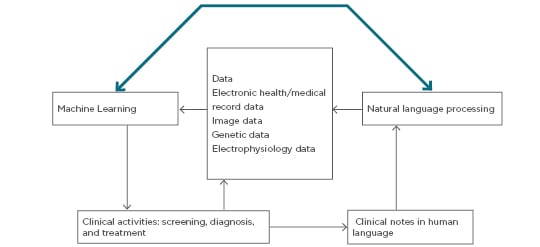
Figure 1: Schematic diagram of clinical decision making using clinical data cues with natural language processing and machine learning analysis.
ML procedures aid in clustering traits or attributes of patients and integrate the information to determine the probability of different disease outcomes. NLP procedures convert text to machine-readable structured data that can then be analysed using ML techniques (Figure 2).11
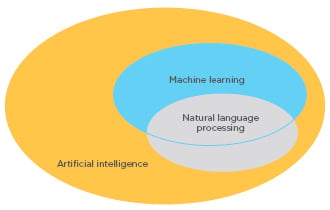
Figure 2: Schematic representation of the relationship between machine learning and natural language processing.
In the entity-attribute relationship, the patient is an entity and the attributes or traits include data such as age, gender, clinical symptoms, past history, diagnosis, biochemical parameters, diagnostic imaging, EP tests, physical examination results, medication, genetic testing, and follow up. Prognostic outcomes are also important in the analysis. Disease markers, disease indicators, quantitative disease levels such as tumour sizes and tumour marker levels, and 5 or 10-year survival rates are also important factors collated in records today.
UNSUPERVISED, SUPERVISED, AND REINFORCEMENT LEARNING
The ML algorithms can be broadly divided into unsupervised learning, supervised learning, and reinforcement learning. Unsupervised learning helps in feature extraction, while supervised learning is used for predictive modelling. Prediction elucidates the relationship between a patient trait and the outcome of interest, which is then used outside the training dataset. The reinforcement learning mode aims at ‘strategic sequencing of rewards and punishments’. Semi-supervised learning is a hybrid system used for scenarios in which the outcome is missing for certain subjects.12,13
Unsupervised Learning Methods
Among the unsupervised learning methods, clustering and principal component analysis (PCA) are used to cluster subjects with similar traits together. Clustering algorithms provides cluster labels for patients, maximising similarity of patients within clusters and minimising similarity of patients between different ones. Hierarchical clustering, k-means clustering, and Gaussian mixture clustering are the most common clustering algorithms used in medicine. PCA is the mainstay of dimension reduction to represent the trait in some important dimensions and ignore background noise from noncontributing dimensions when the trait is recorded in a large number of dimensions. This analysis can be understood as the effort to project the trait in fewer principal component directions, such as the number of genes in a genome-wide association study;12-15 however, when too many dimensions are used in the analysis the random effects interfere with one another and cause significant issues.13,14
Supervised Learning Methods
In supervised learning, the outcomes of the subjects are studied depending on their traits using a particular ‘training process’ to determine a function or an algorithm that correlates or associates outputs with the traits that are closest to the outcomes. Supervised learning has been used to study and provide information about clinically relevant results and therefore is one of the most commonly used AI techniques in medicine.16 Unsupervised learning is often used in the preprocessing step for dimensionality reduction or cluster recognition to make supervised learning more data-efficient. Once a more powerful AI becomes available, this step will be automatically transferred from one dimension to another whether related or unrelated. In the present day, many are conversant around the techniques used for medical applications, including linear regression, logistic regression, naïve Bayes, decision tree, nearest neighbour, random forest, discriminant analysis, support vector machine (SVM), and neural network.16
Support Vector Machine
SVM and neural networks are the most popular techniques used because most of the work is related to image, genetic, and EP analysis. SVM and neural networks have been extensively used in oncological, neurological, and cardiovascular diseases.16 SVM has been used for early detection of Alzheimer’s disease, identification of imaging biomarkers for neurological and psychiatric diseases, and diagnosis of cancer. Model parameter determination, a convex optimisation problem, is always the global optimum in SVM. Mathematically, SVM is a discriminative classifier defining a separating hyperplane between two or more groups. It is a form of supervised learning using labelled training data where the algorithm creates an optimal hyperplane. In a two-dimensional space, the hyperplane can be represented by a line dividing the plane into two parts with different classes on either side of the line.15,17
SVM is therefore extensively employed in medical research. It classifies the subjects into two or more groups; the outcome Yi is a classifier. It operates on the assumption of mutually exclusive groups, meaning that subjects can be separated into two or more groups through decision boundaries defined by the traits. A weighting is applied to the trait depending on its relative importance on affecting the outcome. The sum total of all weights should ideally add up to 1.0 or 100.0%. The goal of this training is to determine the optimal weight so that the outcomes are explained as much as possible in the resulting classification by the traits. The ideal goal is to minimise the error of classifying a patient to the wrong group based on outcome (Yi), also called misclassification error. The outcome is Yi and the tuning parameters are used to draw a plane that separates the groups in nonprobabilistic space. This can be done using many methods, but the most common ones still retain minimisation of quadratic loss function or ordinary least squares.15 The main tuning parameters used are kernel, regularisation, gamma, and margin. The learning of the hyperplane in linear SVM involves transformation of the problem using a linear equation. For linear kernel, the equation for prediction for a new input using the dot product between the input vector (x) and each support vector (xi) is calculated by f(x)= B(0) + sum[ai* (x,xi)].
This equation involves the calculation of inner products of a new input vector (x) with all support vectors from the training data incorporated. Coefficients B0 and ai need to be calculated for each input from the training data by the learning algorithm. The polynomial kernel is calculated by K(x,xi)= 1 + sum(x*xi)^d and exponential as K(x,xi)= exp(-gamma*sum((x — xi²)). Regularisation or C parameters reflect the effort to avoid misclassifying each training example. Larger values of C will choose a hyperplane that will be better at getting all the training points classified correctly even if that line or plane has to curve repeatedly. A small C value makes the optimiser define a larger margin separating the hyperplane at the cost of misclassifying more points. A margin is a separation of line from the closest class points; a good margin is one where this separation is larger for both the classes. Gamma parameter shows how far the influence of a single training example reaches. A low gamma reflects ‘far’ points, away from the plausible separation line that is considered in the calculation, and high values mean ‘close’ points; therefore, points close to the plausible separation line are considered. SVM has been used in the diagnosis of cancer imaging biomarkers18 of neurological and psychiatric disease.17,19
Neural Network
Neural network is a technique in which the association between the input variables and the outcome are represented by multiple hidden layer combinations of prespecified functions. The goal of the equation is to estimate the weights through input and outcome data so that the average error between the outcome and their predictions is minimised. Neural networks have been used to diagnose stroke, to diagnose cancer from 6,567 genes,20 to predict breast cancer using texture information from mammographic images,21 and to diagnose Parkinson’s disease from motor and nonmotor symptoms and neuroimages.22
DEEP LEARNING
Deep learning is a neural network with multiple hidden layers and is capable of exploring complex nonlinear data patterns and high volumes of data. Increased data volumes and complexities have ensured the increasing popularity of deep learning. In 2016, deep learning achieved a 100% year on year increase in the field of medical research. Deep learning is extensively used in imaging analysis because the images are complex and the number of pixels generated is high. Commonly used deep learning algorithms in medicine include recurrent neural network, deep belief network, deep neural network, and convolution neural network (CNN), of which remains the most popular.15,23
Classical ML algorithms have failed to handle high-dimensionality data with large numbers of traits. The images contained thousands of pixels as traits and hence increased dimensionality. Initial solutions depended on dimension reduction by using a subset of pixels as features and later performing the ML algorithms, but information loss in images can be substantial with such handling. Unsupervised learning techniques like PCA or clustering have been used for data driven dimension reduction.15,23 Training datasets with experts performing heuristic analysis was an alternative approach used in radiology and ophthalmology, with CNN being used for high-dimensional image analysis.24 Normalised pixel values were used from images and weighted in convolution layers with iterative sampling in the subsampling layers. A recursive function of weighted input values gave the final outputs. The weights were optimised to minimise the average error between outcomes and predictions24 and the popular software packages Caffe, TensorFlow, and Cognitive Toolkit provided CNN support.25-27
CNN has been used for disease diagnosis in congenital cataract disease using the ocular images,28 in diagnosing skin cancer from clinical images,29 and for detecting referable DR through the retinal fundus photographs.30 The sensitivity and specificity of the algorithms were >90%, and the CNN performance was comparable to experienced physicians in the accuracy for classifying both normal and disease cases. In 2009, Retinopathy Online Challenge used competition fundus photographic sets from 17,877 patients with diabetes who had not previously been diagnosed with DR, consisting of two fundus images from each eye. It is universally agreed that a combination of blood vessel parameters, microaneurysm detection, exudates, texture, and distance between the exudates and fovea are among the most important features to detect the different stages of DR.31 Nayak et al.32 used the area of the exudates, blood vessels, and texture parameters, analysed through neural networks, to classify the fundus image into normal, nonproliferative DR, and proliferative DR. A detection accuracy of 93% with a sensitivity of 90% and a specificity of 100% were reported. SVM classified fundus images into normal, mild, moderate, severe, and prolific DR classes, with a detection accuracy of 82%, sensitivity of 82%, and specificity of 88%. Lee et al.33 described a software to grade the severity of haemorrhages and microaneurysms, hard exudates, and cotton-wool spots of DR as a means to classify the disease.33
Clinical information in narrative text, physical examination, clinical signs, laboratory reports, operative notes, and discharge summaries are unstructured and not understood by computer programs. NLP is important in such situations because it parses the narrative text to extract information to assist the clinician in decision making.14 The NLP pipeline has text processing and classification components. Using text processing, a series of disease-relevant keywords are identified in the clinical notes in databases. A subset of the keywords, through domain reduction, are selected and validated to analyse the structured data to aid clinicians in the decision-making process.12,13
The NLP pipelines have been used to read chest X-ray reports to alert physicians regarding the need for anti-infective therapy,34 to monitor laboratory-based adverse effects35 and variables associated with cerebral aneurysms disease,36 and to extract cases of peripheral vascular disease from narrative clinical notes37 with a high degree of accuracy.
Real-life implementation of AI technologies is still in nascent stages in medicine. It is facing hurdles including regulation, the interoperability of information algorithms, and availability of data for algorithms especially for validation after the training is over. Lack of standards to assess the safety and efficacy of AI systems has resulted in a proliferation of proprietary algorithms which do not crosstalk. The U.S. Food and Drug Administration (FDA) has recently advised classifying AI systems as ‘general wellness products’ putting them under loose regulation. Furthermore, rules for adaptive design in clinical trials were formulated for the assessment of operating characteristics of AI systems.38
There are no explicit laws covering data transfer for processing in many countries. Extraordinarily large amounts of data can be processed by an indirect third-party by service providers, recoding data according to USA laws, such as the deal between Google DeepMind and the Royal Free London NHS Foundation Trust which lead to debate in 2017. That agreement was criticised on the grounds of violating the Caldicott principles by transferring more data than necessary and blurring the line between the data controllers and data processors. Direct care providers need to be careful when sharing data with third-parties who are not in a direct care relationship with the patient in question. If explicit consent and notice have not been given then all deidentified data, whether labelled or unlabelled, should come into public domain and be published by a statutory body. This will keep a check on illegal proprietary exploitation of the data and force the data processor to seek limited amounts of data for exchange.39 Since, in the absence of consent, such deidentified datasets should be considered community resource, there is logic in placing it in the hands of the community, thereby enabling policing of these data exchanges is an extremely important issue that the governments across the world must consider seriously. With transfer learning and AGI, humans may not even be able to understand how the machine handled the data in the AI tools employed by them. Therefore, there is an extreme need to ensure the formulation of and adherence to principles of computational bioethics.39 Once the AI system is deployed there should be a lack of continued data supply for the development and improvement of the system, and after initial training with historical data, little fine tuning and further algorithm development should be done. Additionally, no incentives for sharing data among users of the system should be made (Figure 3).
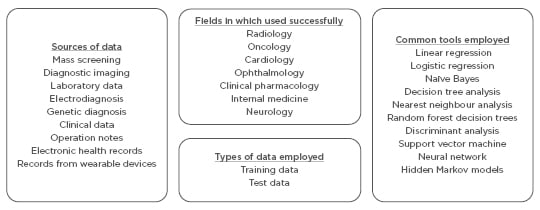
Figure 3: Overview of artificial intelligence in healthcare and challenges in obtaining and using data.
Artificial intelligence systems require continuous training using data from clinical studies.
The data employed can broadly be classified into training data and test data.13,14
WHO PAYS FOR FINANCING THESE SYSTEMS?
Tax incentives have been offered, and the more recent emergence of the insurance companies as primary payers has started to shift the paradigm from ‘payments based on treatment volume’ to ‘payments based on the treatment outcomes and fiscal efficiency’. Thus, the treating physicians, pharmaceutical companies, and patients have greater incentives to compile and exchange information.40
Development of complex algorithms is laborious and requires skilled resources. Once developed, they can be used in simple devices with standard hardware, similar to how face recognition was implemented on mobile phones. These algorithms can bring down the costs of the screening programmes immensely. Though in its nascent stages, the potential of this deep learning for use in DR screening programmes has been recognised and the latest results from Google’s recent attempt are very encouraging.39
FUTURE CONCERNS AND COMPENSATION OF THE CONTRIBUTORS
It must be remembered that these algorithms are being developed using resources from the communities at large and these communities must be suitably compensated. The ownership of the pooled data also needs to be looked at more carefully, especially with the increased use of wearable technologies. The proprietary platforms and lack of data interexchange, as well as absence of crosstalk, was a major inhibiting factor in the development of such technologies. The medical personnel need to become guides in the development of bioethics-based rules. It is important that adequate care is given to address all these bioethical concerns, because these rules will form the basis for all future developments and documentation. The supremacy of the human will should not become subservient to AI, even after it subjectively becomes increasingly more intelligent than the human beings. Clinical research, with the use of emerging tools, expanding volumes of data, structured databases, and rich information is now answering ever more complicated clinical questions and allowing forward chaining to real-life clinical questions. Prevention, diagnosis, treatment, and prognosis are becoming better defined with AI techniques, but the onus of ensuring that the machines do not run amok in the future lies with the clinicians and society at large.
CONCLUSIONS AND RECOMMENDATIONS
AI applications in healthcare have tremendous potential for being useful; however, success is dependent on clean, high-quality healthcare data, which requires impeccable planning and execution. Data capture, storing, preparation, and mining are critical steps and a standardised clinical vocabulary and the sharing of data across platforms is required. Bioethical standards in collecting and using data cannot be overemphasised. The authors hope that this paper makes the stakeholders realise their responsibility, contributes to AI in healthcare literature, and guides the development of tools to be employed for practice.








