Understanding the interplay between the immune system and gastrointestinal tract (GIT) microbiota is an evolving area of research. Emerging evidence suggests that GIT microbiota play a key role in the development of innate and adaptive immunity,1,2 and may induce or perpetuate aberrant immune function in autoimmune diseases, such as inflammatory bowel disease3 and rheumatoid arthritis.4
Systemic sclerosis (SSc) is a devastating autoimmune disease, which profoundly affects GIT function. While the cause of GIT dysfunction in SSc is unknown, our group recently characterised the lower GIT microbiota in a small, unselected cohort of patients with SSc.5 We found significant microbial community differences in SSc patients versus healthy controls, in the caecum and sigmoid regions, in patients undergoing colonoscopy.5 We also observed significant genus-level differences between SSc patients and healthy controls, including decreased beneficial commensal genera such as Faecalibacterium, Clostridium, and Rikenella, as well as increased potentially pathobiont genera, including Fusobacterium, Prevotella, Ruminococcus, Akkermansia, and the uncommon γ-Proteobacteria, Erwinia and Trabulsiella.5
Our subsequent study, analysing faecal samples of SSc patients in two independent SSc cohorts, identified similar microbial community differences in SSc patients and healthy controls (Table 1).6 Furthermore, we identified specific microbial genera associated with GIT symptom severity.6
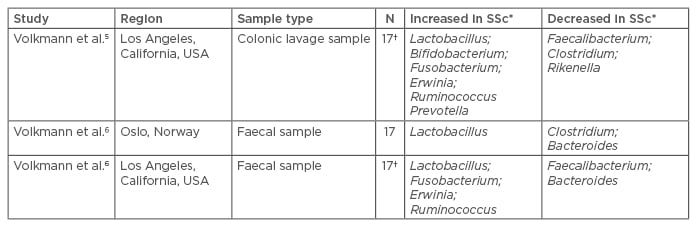
Table 1: Summary of increased and decreased microbial taxa in SSc patients versus controls.
*Relative to healthy controls.
†These are the same SSc subjects. In Volkmann et al.,5 we collected lavage specimens from the cecum and sigmoid colon during colonoscopy; in Volkmann et al.,6 we collected faecal specimens. However, the healthy control groups comprised different individuals in the two studies.5,6 SSc: systemic sclerosis.
However, these studies were cross-sectional; therefore, it is unclear whether the relationships observed between specific genera and GIT symptoms are causational and/or persist with time. To address this limitation, in the present study, we collected faecal specimens and measured GIT symptom severity in SSc patients every 3 months, over a 12-month period. The microbiota from the stool specimens were profiled by multiplex sequencing for bacterial rRNA genes, using an Illumina HiSeq 2500 (Illumina Inc., San Diego, California, USA) sequencing technique. All samples were analysed simultaneously to avoid any batch effects. We assessed GIT symptoms using the GIT 2.0, a valid measure of GIT symptom severity in SSc patients.7 The questionnaire consists of seven domains, and has been translated and validated in several languages. Scores on the GIT 2.0 can indicate self-rated severity (i.e. none/mild versus moderate versus severe/very severe disease) of GIT involvement based on previously published score thresholds.7
We discovered that the absolute and relative abundances of specific genera did not significantly change within individual SSc patients over a 12-month period. In addition, GIT 2.0 scores did not significantly change over the course of the study within each SSc patient. However, we did find that patients with longer disease durations had increased GIT symptoms over time. In addition, patients who had a lower abundance of Bacteroides throughout the study had increased GIT symptoms over time, even after controlling for age, sex, ethnicity, disease duration, and SSc subtype (i.e. limited versus diffuse cutaneous disease).
These findings provide further evidence that specific microbial genera may contribute to the GIT phenotype in SSc. Furthermore, these results suggest that efforts to replete these key genera (e.g. supplemental probiotics, faecal transplantation) may help to restore GIT microbiota homeostasis and potentially improve symptoms in SSc.
The present research likely represents the tip of the iceberg in SSc microbiome research. Future research efforts in this area are greatly needed to better understand the relationships between GIT microbiota, immune responses, and clinical outcomes in SSc.